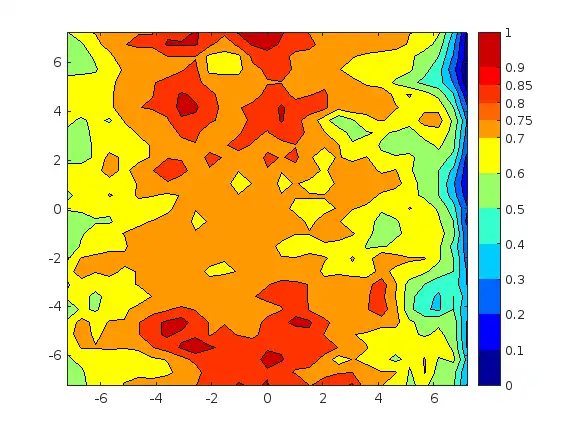
How to change the scale of color bar to discrete ?


hi, I have my contour plot like this figure, I want to change the scale of colorbar to a discrete one with the scale value [ 0, 0.1 ,0.2 ,0.3 ,0.4 ,0.5 ,0.6 ,0.7 ,0.75 ,0.8 ,0.85, 0.9 ,1] how to do it?
 John Williams answered .
2025-11-20
John Williams answered .
2025-11-20
x = [-7.22 -6.70 -6.19 -5.67 -5.15 -4.64 -4.12 -3.60 -3.09 -2.56 -2.04 -1.53 -1.00 -0.51 0.02 0.54 1.06 1.58 2.10 2.60 3.12 3.64 4.16 4.69 5.18 5.70 6.23 6.75 7.25];
y = [7.20 6.70 6.18 5.68 5.14 4.62 4.11 3.59 3.09 2.57 2.05 1.54 1.02 0.51 -0.01 -0.53 -1.03 -1.55 -2.07 -2.59 -3.09 -3.61 -4.14 -4.65 -5.16 -5.67 -6.19 -6.71 -7.22];
strain_12 = [0.48 0.64 0.66 0.7 0.68 0.72 0.76 0.78 0.9 0.88 0.86 0.82 0.88 0.84 0.9 0.88 0.9 0.86 0.88 0.82 0.82 0.8 0.8 0.78 0.76 0.72 0.7 0.68 0.64 0.5 0.64 0.66 0.76 0.68 0.76 0.76 0.78 0.82 0.82 0.8 0.8 0.82 0.84 0.88 0.84 0.86 0.84 0.86 0.8 0.78 0.74 0.72 0.74 0.74 0.72 0.72 0.68 0.64 0.54 0.7 0.68 0.76 0.7 0.68 0.7 0.74 0.76 0.72 0.8 0.86 0.84 0.9 0.92 0.86 0.88 0.86 0.86 0.84 0.8 0.76 0.74 0.7 0.7 0.74 0.72 0.7 0.68 0.48 0.68 0.72 0.74 0.72 0.7 0.7 0.76 0.82 0.82 0.86 0.88 0.84 0.88 0.88 0.88 0.86 0.88 0.9 0.94 0.9 0.86 0.82 0.78 0.76 0.76 0.76 0.76 0.7 0.48 0.66 0.7 0.68 0.7 0.72 0.7 0.74 0.76 0.76 0.82 0.88 0.84 0.84 0.86 0.8 0.82 0.84 0.82 0.88 0.88 0.9 0.88 0.82 0.76 0.78 0.74 0.8 0.72 0.48 0.66 0.68 0.66 0.7 0.78 0.8 0.82 0.82 0.8 0.84 0.9 0.92 0.86 0.84 0.8 0.82 0.82 0.82 0.86 0.92 0.9 0.88 0.82 0.76 0.76 0.74 0.78 0.7 0.5 0.62 0.54 0.56 0.64 0.78 0.86 0.82 0.82 0.82 0.78 0.82 0.86 0.82 0.8 0.82 0.82 0.82 0.8 0.82 0.84 0.82 0.78 0.78 0.74 0.72 0.68 0.7 0.64 0.52 0.62 0.54 0.58 0.64 0.76 0.84 0.82 0.8 0.82 0.8 0.82 0.84 0.84 0.84 0.82 0.8 0.78 0.76 0.76 0.76 0.76 0.76 0.72 0.74 0.7 0.68 0.7 0.7 0.48 0.6 0.58 0.62 0.66 0.76 0.86 0.8 0.78 0.8 0.78 0.82 0.84 0.84 0.82 0.82 0.8 0.8 0.78 0.76 0.78 0.78 0.8 0.76 0.74 0.72 0.7 0.72 0.68 0.48 0.62 0.66 0.7 0.74 0.74 0.78 0.72 0.74 0.74 0.74 0.8 0.76 0.78 0.78 0.8 0.76 0.74 0.74 0.78 0.78 0.8 0.78 0.76 0.74 0.78 0.76 0.8 0.72 0.46 0.62 0.7 0.76 0.76 0.78 0.82 0.72 0.76 0.74 0.76 0.78 0.76 0.76 0.82 0.82 0.78 0.76 0.78 0.82 0.78 0.8 0.78 0.8 0.78 0.78 0.78 0.76 0.68 0.44 0.6 0.7 0.72 0.7 0.7 0.68 0.68 0.74 0.72 0.74 0.72 0.72 0.74 0.78 0.82 0.82 0.82 0.8 0.82 0.78 0.82 0.78 0.8 0.74 0.74 0.7 0.7 0.62 0.44 0.64 0.72 0.7 0.7 0.68 0.64 0.72 0.76 0.8 0.8 0.76 0.76 0.78 0.78 0.8 0.8 0.78 0.76 0.76 0.78 0.8 0.78 0.78 0.76 0.72 0.72 0.68 0.64 0.4 0.62 0.7 0.72 0.72 0.68 0.66 0.72 0.72 0.76 0.78 0.74 0.78 0.82 0.8 0.8 0.8 0.78 0.76 0.74 0.8 0.78 0.76 0.74 0.72 0.68 0.68 0.66 0.64 0.42 0.58 0.68 0.72 0.76 0.72 0.7 0.74 0.76 0.76 0.74 0.74 0.74 0.78 0.76 0.78 0.78 0.8 0.78 0.76 0.78 0.74 0.72 0.74 0.7 0.7 0.72 0.7 0.7 0.38 0.52 0.62 0.68 0.7 0.72 0.74 0.8 0.8 0.82 0.76 0.76 0.76 0.76 0.8 0.78 0.76 0.78 0.78 0.76 0.76 0.74 0.76 0.76 0.74 0.76 0.72 0.7 0.68 0.36 0.5 0.62 0.66 0.72 0.78 0.8 0.82 0.78 0.78 0.7 0.74 0.76 0.74 0.8 0.76 0.72 0.78 0.78 0.78 0.8 0.82 0.78 0.76 0.72 0.74 0.72 0.72 0.7 0.38 0.54 0.62 0.64 0.7 0.76 0.74 0.8 0.8 0.8 0.74 0.8 0.8 0.76 0.82 0.76 0.76 0.8 0.82 0.8 0.84 0.86 0.82 0.78 0.74 0.78 0.74 0.72 0.72 0.4 0.6 0.66 0.66 0.7 0.7 0.7 0.76 0.74 0.76 0.74 0.76 0.84 0.82 0.84 0.78 0.8 0.82 0.84 0.8 0.82 0.82 0.76 0.74 0.74 0.78 0.7 0.66 0.66 0.44 0.64 0.7 0.66 0.66 0.64 0.68 0.72 0.7 0.76 0.76 0.78 0.82 0.8 0.8 0.82 0.84 0.82 0.8 0.82 0.8 0.8 0.76 0.74 0.74 0.72 0.7 0.66 0.66 0.46 0.66 0.74 0.72 0.66 0.68 0.7 0.68 0.66 0.72 0.76 0.8 0.86 0.86 0.82 0.84 0.82 0.78 0.78 0.84 0.84 0.78 0.78 0.76 0.74 0.72 0.72 0.64 0.68 0.48 0.68 0.78 0.78 0.72 0.74 0.76 0.74 0.68 0.64 0.72 0.8 0.84 0.9 0.84 0.84 0.8 0.8 0.84 0.88 0.88 0.82 0.76 0.78 0.76 0.68 0.7 0.68 0.7 0.42 0.62 0.74 0.74 0.74 0.76 0.78 0.78 0.8 0.78 0.84 0.86 0.86 0.9 0.86 0.84 0.82 0.8 0.86 0.9 0.92 0.84 0.84 0.82 0.8 0.74 0.74 0.72 0.72 0.36 0.56 0.66 0.72 0.76 0.74 0.8 0.8 0.8 0.78 0.84 0.82 0.8 0.86 0.84 0.8 0.8 0.8 0.84 0.88 0.9 0.86 0.82 0.82 0.8 0.74 0.72 0.72 0.72 0.3 0.48 0.58 0.6 0.64 0.68 0.72 0.74 0.78 0.8 0.8 0.78 0.76 0.82 0.82 0.78 0.78 0.78 0.8 0.84 0.86 0.84 0.82 0.78 0.78 0.74 0.72 0.74 0.72 0.28 0.44 0.58 0.64 0.7 0.72 0.72 0.7 0.72 0.76 0.76 0.82 0.8 0.88 0.84 0.8 0.74 0.74 0.74 0.84 0.82 0.8 0.8 0.8 0.76 0.74 0.68 0.66 0.66 0.32 0.48 0.62 0.66 0.7 0.7 0.72 0.74 0.76 0.76 0.76 0.8 0.8 0.86 0.86 0.84 0.74 0.74 0.74 0.82 0.82 0.82 0.78 0.78 0.8 0.74 0.7 0.66 0.68 0.32 0.5 0.66 0.72 0.74 0.8 0.8 0.8 0.8 0.78 0.8 0.84 0.86 0.88 0.92 0.9 0.84 0.78 0.84 0.9 0.9 0.9 0.84 0.84 0.82 0.78 0.74 0.68 0.66 0.34 0.54 0.7 0.74 0.76 0.8 0.78 0.78 0.8 0.78 0.8 0.82 0.84 0.9 0.96 0.96 0.9 0.84 0.86 0.9 0.86 0.88 0.82 0.82 0.82 0.76 0.76 0.74 0.68];
[a,b] = meshgrid(x,y);
gama_12 = reshape(strain_12,29,29);
shear = rescale(gama_12,0,1);
contourf(b,a,shear);
lvls = [ 0, 0.1 ,0.2 ,0.3 ,0.4 ,0.5 ,0.6 ,0.7 ,0.75 ,0.8 ,0.85, 0.9 ,1];
% find the indices where the level difference is "big" (0.1):
lvl_idx = find(diff(lvls) > 0.075); % (closer to 0.1 than to 0.05)
n_big = numel(lvl_idx);
% create a colormap with an extra color for each big level difference:
cmap = jet(numel(lvls)-1+n_big);
% change indices in lvls to indices in the colormap:
cmap_idx = lvl_idx;
for ii = 1:n_big
cmap_idx(ii) = lvl_idx(ii) + nnz(lvl_idx <= lvl_idx(ii));
end
% duplicate the colors in the colormap at the big level indices:
cmap(cmap_idx,:) = cmap(cmap_idx-1,:);
% apply the colormap, create the colorbar, and set the ticks to lvls:
colormap(cmap);
cb = colorbar();
cb.Ticks = lvls;